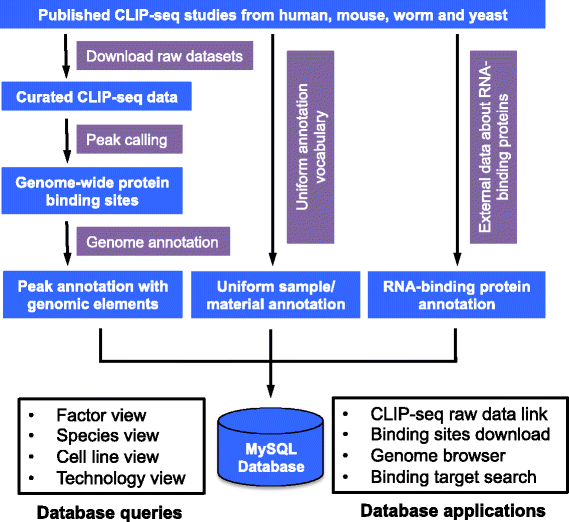
Main steps of the bioinformatics workflow to analyze CLIP-seq data with... | Download Scientific Diagram

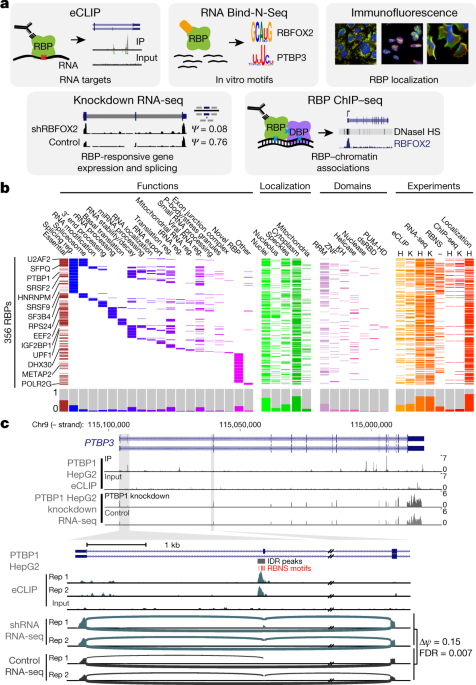
Global Approaches in Studying RNA-Binding Protein Interaction Networks: Trends in Biochemical Sciences
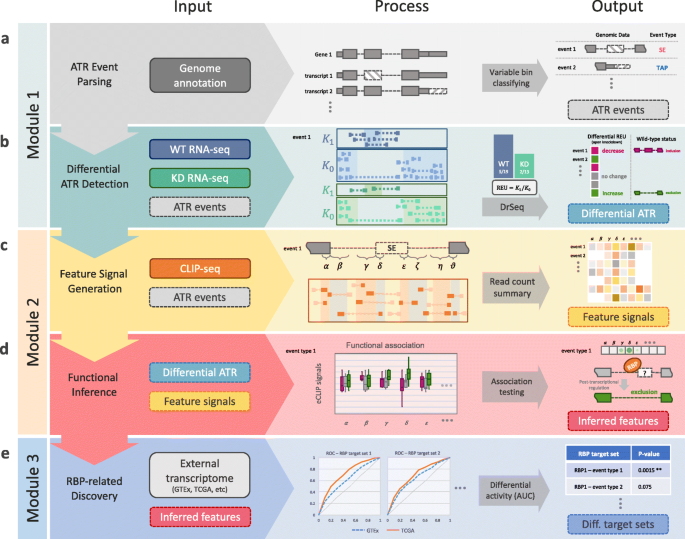
SURF: integrative analysis of a compendium of RNA-seq and CLIP-seq datasets highlights complex governing of alternative transcriptional regulation by RNA-binding proteins | Genome Biology | Full Text

Computational approaches for the analysis of RNA–protein interactions: A primer for biologists - Journal of Biological Chemistry

Figure 5 from CLIP-seq analysis of multi-mapped reads discovers novel functional RNA regulatory sites in the human transcriptome | Semantic Scholar
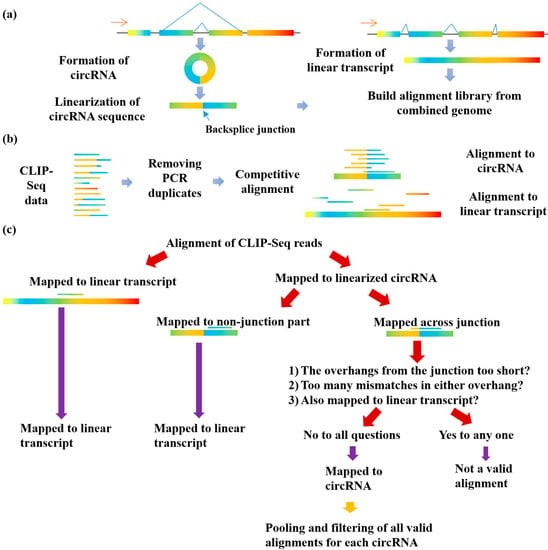
Genes | Free Full-Text | Large-Scale Profiling of RBP-circRNA Interactions from Public CLIP-Seq Datasets | HTML

Interpretation of the AGO-binding model learned from CLIP-seq data. (A)... | Download Scientific Diagram

SURF: integrative analysis of a compendium of RNA-seq and CLIP-seq datasets highlights complex governing of alternative transcriptional regulation by RNA-binding proteins | Genome Biology | Full Text



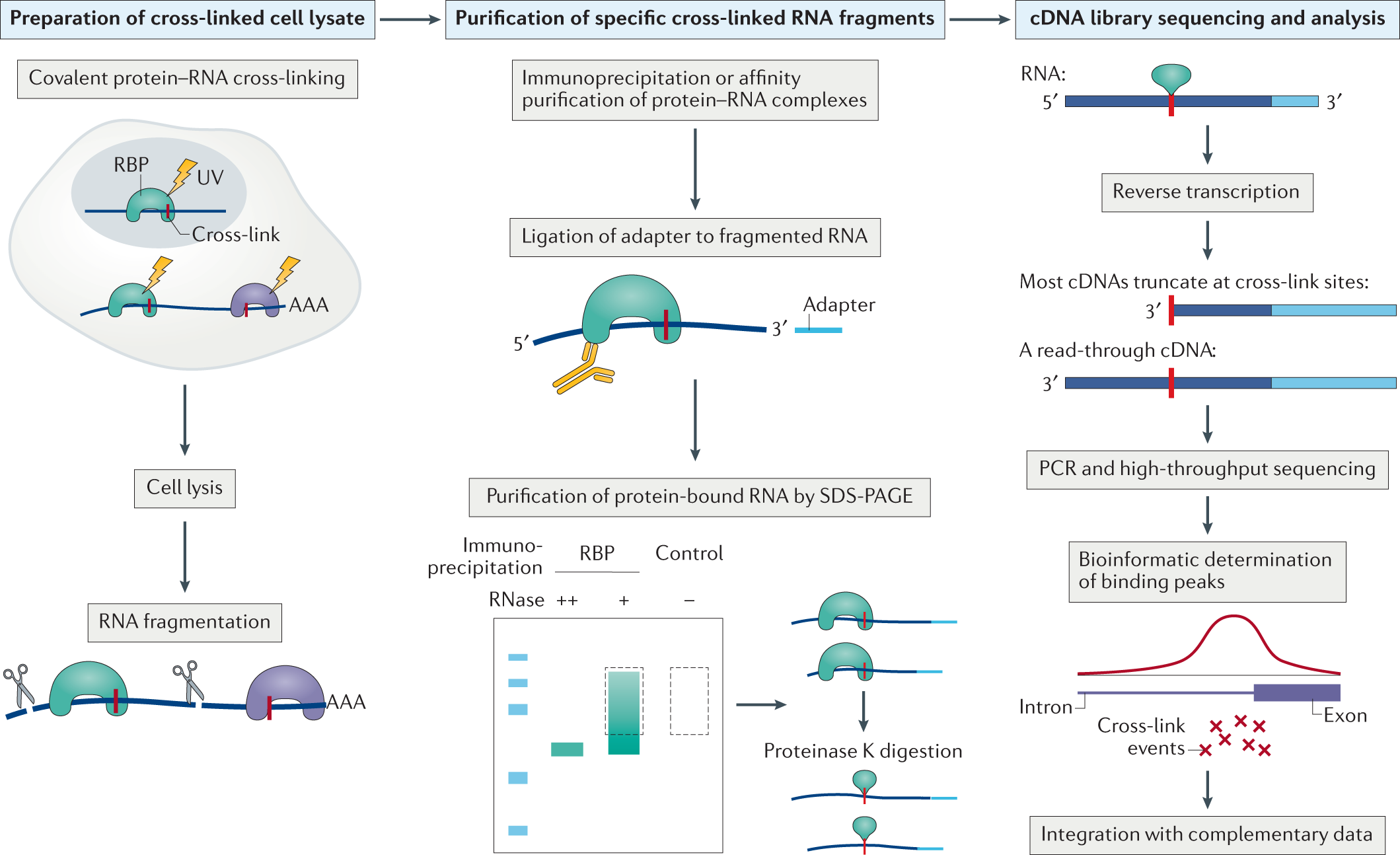
![PDF] Computational analysis of CLIP-seq data. | Semantic Scholar PDF] Computational analysis of CLIP-seq data. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/6e732e2e32a7bb333de34a0a4696eac4ddf3e45d/14-Figure3-1.png)